|
2/2/2024 0 Comments 10x chromiumSingle nucleus ATAC Sequencing – This kit is used to generate ATAC-seq libraries for measurement of chromatin accessibility at single-nucleus resolution. A CRISPR/sgRNA screening module compatible with the 5′ Gene Expression kit is coming soon. Like the 3′ Gene Expression kit, additional add-on modules are available for cell surface protein expression and sample multiplexing. Currently, 10X Genomics only sells these modules for human and mouse V(D)J amplification. 
The main reason to choose this kit over the 3′ Gene Expression kit is the immune repertoire profiling add-on module, which allows for the parallel PCR enrichment and library preparation of B cell/T cell receptor V(D)J sequences. Single cell 5′ Gene Expression/Immune Profiling – This kit also generates single cell RNA-seq libraries through capture at the 5′ end by capturing the TSO sequence added to this end of the transcripts in a template-switching reverse transcription reaction. Add-on modules are available for cell surface protein expression (analogous to CITE-seq), CRISPR/sgRNA screening, and sample multiplexing. The kit employs polyA-based capture of mRNA at the 3′ end to generate dual indexed libraries containing both a cell barcode identifying the cell of origin as well as a unique molecular identifier (UMI), which will be unique to every transcript captured. Single cell/single nucleus 3′ Gene Expression– The standard “workhorse” kit for single cell/nucleus RNA sequencing. If this happens, we will keep in touch with you to let you know when we receive the kits, so that you are not bringing over samples on a day when we do not have the reagents to process them. While 10X Genomics is usually able to deliver kits a day or two after they are ordered, they do suffer from occasional back-order issues. We ask that you provide a funding string at the time we place the order so that we can eventually bill for the cost of the kits even if your project plans do not end up proceeding. For any other assay, we will order the kits specifically for your project. Nuclei suspension can also be prepared and sequenced.Ī note about these kits: We only maintain a stock of the 3′ Gene Expression kits.70,000 reads per cell is appropriate for expression analysis. 50,000 reads per cell should be sequenced for saturating gene detection.Captures approximately 60% of loaded cells.10x Chromium instrument capable of capturing 500 to 10,000 cells per sample up to 80,000 cells per chip.Samples are barcoded individually for a multiplexed sequencing reaction. 8-channel microfluidic chip allowing up to 8 samples to be processed simultaneously.Please e-mail schedule a meeting to discuss your proposed experiment and timeline! 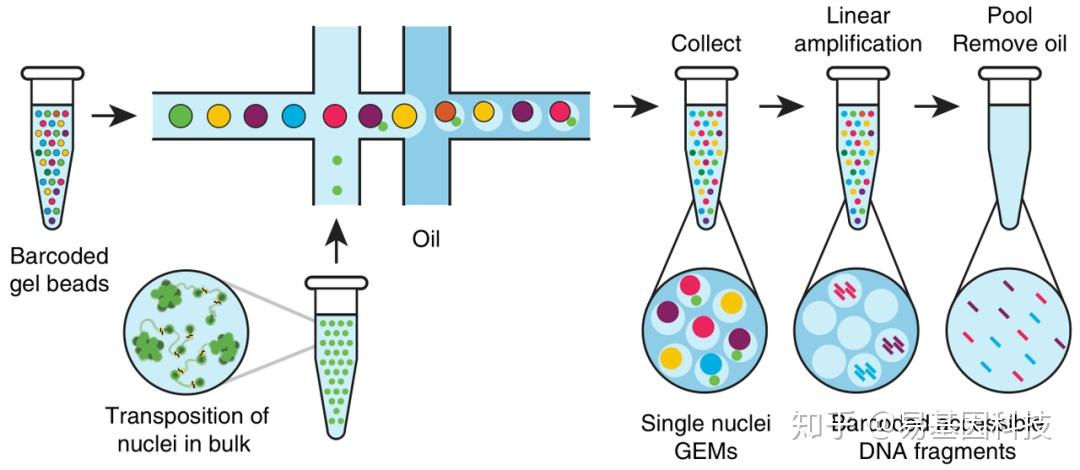
The resulting data can then by analyzed and explored using the freely downloadable Loupe Cell Browser software ( link for Loupe Cell Browser free download, tutorial, and sample data sets).Īdditionally, 10x Genomics offers assays for single cell immune profiling (sc-V(D)J enrichment kit), single cell copy number variation profiling, single cell chromatin accessibility analysis (sc-ATAC-Seq), de novo assembly, and genome or exome sequencing. User provided single cell or single nuclei suspensions are loaded onto the Chromium iX to generate single cell barcoded gel-bead emulsions from which Illumina-compatible library preps are generated and sequenced on the NovaSeq in collaboration with UWBC DNA Sequencing Facility followed by optional analysis using a custom developed single cell data analysis pipeline by the UWBC Bioinformatics Resource Center. The Single Cell Gene Expression Assay is able to generate single cell transcriptomic measurements from up to 10,000 cells per cell suspension with up to 65% capture rate efficiency enabling the identification of rare cell types from heterogeneous populations. 
The 10x Genomics Chromium platform combines microfluidics with molecular barcoding to provide mRNA quantification from thousands of individual single cells.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
